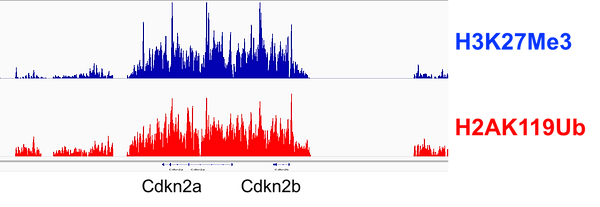
Investigating the epigenetic landscape of different epidermal sub-lineages
During development, a single layer of epidermal progenitors gives rise to the different epidermal sub-lineages. This includes the basal cells that separate the dermis from the epidermis, the suprabasal layers (differentiated cells that constitute the impermeable barrier of the skin), and the Merkel cells, important mechanoreceptors involved in touch sensitivity. During his postdoc, Julian has been interested in exploring the epigenetic changes associated with the establishment of the different epidermal sub-lineages. To this end, he uses transgenic mice to isolate the different epidermal sub-populations by FACS. For example, he utilizes a mouse strain that expresses RFP under the basal cell promoter Krt14, and GFP under the suprabasal promoter Krt10 to isolate basal and suprabasal cells. To isolate Merkel cells, Julian uses Atoh1-GFP reporter mice. To characterize the different histone modifications associated with promoters, enhancers, and repressed and active genes in each of these epidermal cell types, he has optimized ChIPseq protocols to be performed in a very low number of FACS-isolated cells. Additionally, he is performing ATACseq on the same FACS-isolated epidermal cell populations to map open chromatin regions and transcription factor binding sites. Integrating RNAseq, ChIPseq, and ATACseq data from the embryonic epidermal progenitors, as well as the basal, suprabasal, and Merkel cells will allow us to characterize the epigenetic landscape of these cells. These studies will provide us with a better understanding of how the different epidermal lineages are defined and how chromatin structure orchestrates cell differentiation and commitment.